For the 2017 Elton Prize, the Editors selected one winning paper and two highly-commended papers. Last month we featured a blog post about prize winner Natalie Clay, and now we are proud to feature a post by highly-commended author Nick Fountain-Jones. Nick is a postdoc with the Department of Veterinary Population Medicine at the University of Minnesota. Read on to hear the #StoryBehindThePaper
Understanding disease transmission is a major challenge in disease ecology. This is particularly true for viruses that infect social groups of wildlife. Contact between individuals and groups are frequent but not all contacts lead to transmission events and understanding what types of contact matters is difficult. Further muddying the waters, even within the same disease, the types of contact that lead to transmission can vary by genetic variants or ‘subtypes’ of that disease. These subtype-specific differences in transmission not only have implications for managing disease in wildlife but also have fundamental consequences for how that disease evolves. But how to detect differences in transmission between virus subtypes in wild populations? This was the problem that I grappled with when I started my postdoc with Meggan Craft at the University of Minnesota.

Under the stewardship of Craig Packer, the University of Minnesota is home to the famous Serengeti Lion Project (SLP) that has unique long-term data from thousands of lions that provided an ideal system to start to explore what contacts matter for different subtypes of feline immunodeficiency virus (FIV). As with HIV, FIV is genetically diverse and the Serengeti lion population is a ‘melting pot’ of FIV. Three different subtypes are found across the population with all lions infected by at least one subtype with some infected by all three. Even though most FIV infected lions live normal lives, previous research led by Jennifer Troyer revealed that lions infected by FIV subtype C are potentially more likely to die young compared to those infected by B. She also speculated that the contacts that led to transmission also varied, but did not have to tools to test this hypothesis. Armed with community phylogenetic tools I had learned during my PhD at the University of Tasmania, started to quantify subtype differences between the more common subtypes B and C using FIV molecular data together with a wealth of information from 16 prides from the SLP database. I found contacts between pride-mates were important for the transmission of both subtypes, but surprisingly within a pride, whilst mother-cub relationships were universally important for transmission, dads played an important role in the transmission of subtype C to their cubs. The mechanism for this is unclear at this stage, yet as father-offspring relationships are less often considered to impact transmission this was an important finding.
Upscaling this to examine if these subtype differences could shape between-pride transmission dynamics remained a challenge. Whilst working on this problem, I was awarded funding from the infectious disease evolution across scales (IDEAS) program for a research exchange with the University of Glasgow with Roman Biek. A solution came during my stay in Glasgow when I met Roman’s postdoc Maude Jacquot who works on disease genetic networks where two individuals that share genetically similar viruses are ‘connected’. In the same way, Maude created networks for the Serengeti lions so that lion prides that had a similar FIV genetic profile were ‘connected’. I then compared networks generated for each subtype and used more community ecology statistical approaches to overlay different types of networks, such as male immigration where two prides are connected if a male immigrated from one pride to another, to see which one was the best fit. Again males were important for the transmission of FIV subtype C with the male immigration network correlating best with the subtype C network as prides far away from each other on the landscape had similar subtype C genetic profiles. Male lions do most of the dispersing and it turns out they take their FIV subtype C with them. In contrast, subtype B showed a totally different pattern with prides neighbouring each other sharing similar subtype B genetic profiles. Females are the guardians of pride territory and defend it from others so this may be evidence for the importance of aggressive interactions between females in the transmission of FIV subtype B. This new approach to untangling transmission between groups was a product of a collaboration made possible via research exchange, which highlights how important exchanges between labs are. More generally, this paper also demonstrates how integrating network, phylogenetic and community ecology tools can be applied to understand disease dynamics.
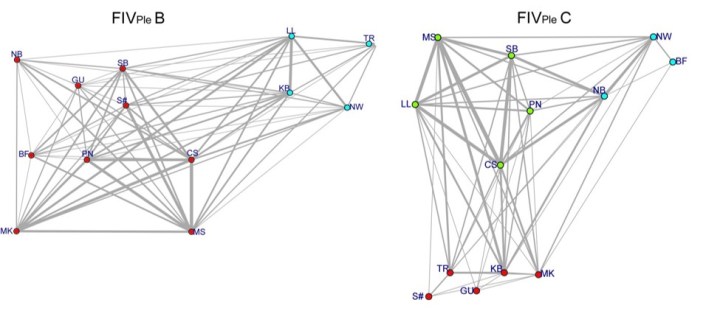
This is just the beginning of understanding on how genetic variation within a disease can have consequences for transmission. Next steps include sampling more of the FIV genome (this study was based on a small part of the FIV genome) from a larger number of individuals to gain more fine scale insight into how FIV spreads in lions. Furthermore, successful transmission of an FIV subtype from lion to lion may also be dependent on the presence (or absence) of other subtypes or other diseases, so exploring this avenue is a logical next step.
More Info:
Fountain‐Jones, N.M. et al. (2017) Linking social and spatial networks to viral community phylogenetics reveals subtype‐specific transmission dynamics in African lions. Journal of Animal Ecology, 86: 1469–1482. https://doi.org/10.1111/1365-2656.12751
