This blog post is provided by Kristy M. Ferraro and Diego Ellis-Soto and tells the #StoryBehindThePaper for their article “A methodological roadmap to quantify animal-vectored spatial ecosystem subsidies“, which was recently published in the Journal of Animal Ecology.
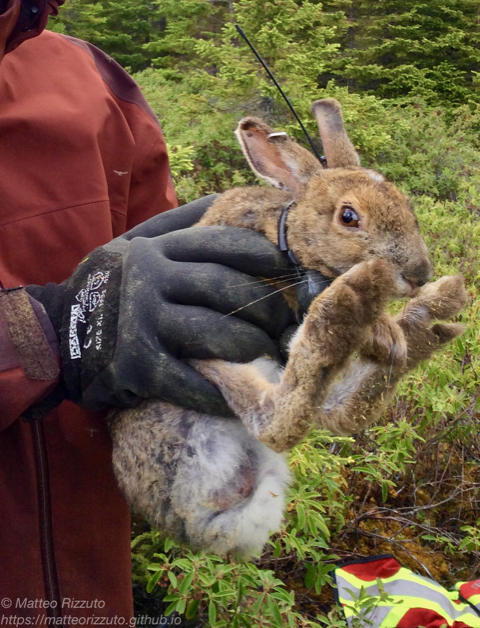
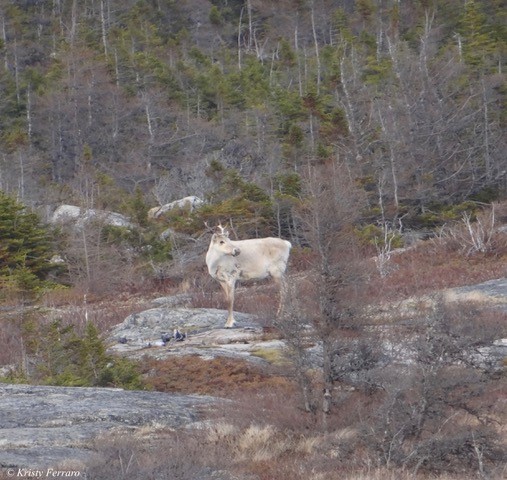
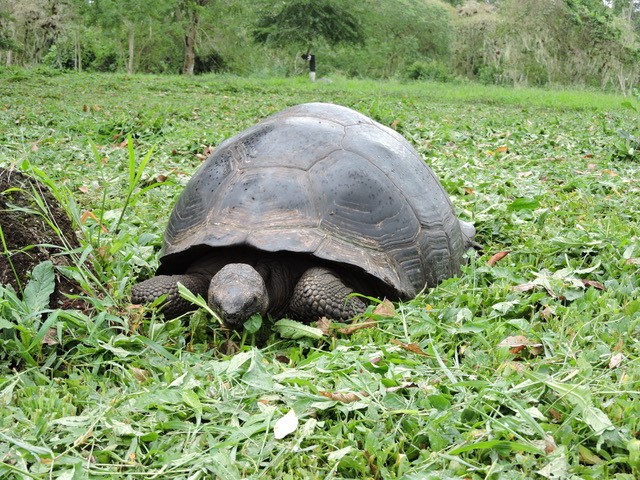
Animals are constantly on the move – whether it’s the snow-shoe hare’s quick hop around its home range or a Galapagos tortoise’s seasonal migration up and down a volcanic island – rarely are animals stagnant in time and space. Yet it’s not just through their tracks that they can connect different ecosystems, through consumption, defecation, and death, animals can move nutrients between systems (Earl & Zollner 2017). As such, animals on the move could be seen as the ‘organismal veins’ that connect ecosystems.
Recently, the number of studies exploring how animals transport nutrients between ecosystems has grown rapidly. Termed zoogeochemistry (Schmitz et al. 2019), this field actively seeks to understand how animals are impacting biogeochemical cycles and providing important ecosystem services through the process of translocating nutrients. These translocations can be extremely important and impactful for systems, providing important subsidies for the receiving ecosystem. For example, through defecation whales in the ocean bring nutrients from the deep ocean to the surface, a redistribution that supports life at the surface of the zone (Roman & McCarthy 2010). Yet as large animal populations and movements are declining globally (Wilcove & Wikelski 2008), untangling the role of these animals is of increasing importance.
Our team has a plethora of study species, including caribou, pumas, vicuñas, snowshoe hares, and giant tortoises. We also come from a range of disciplines, including ecosystem ecology, animal ecology, biogeochemistry, and remote sensing. With a common ground in zoogeochemistry, we began meeting and discussing the importance of untangling the role of animals in the nutrient cycles of their respective ecosystems. Using the conceptual framework of meta-ecosystem theory, which suggests that ecosystems are not necessarily closed but can be connected, we began to dream up questions to explore how animals connect systems through the movement of nutrients. Yet as we sat around a table (this was pre-covid times!), we all wondered the same question: studies of animal-mediated nutrient translocation are logistically difficult, so how can we best harness the tools available to us to study the zoogeochemical effects of our vastly different study systems and species?
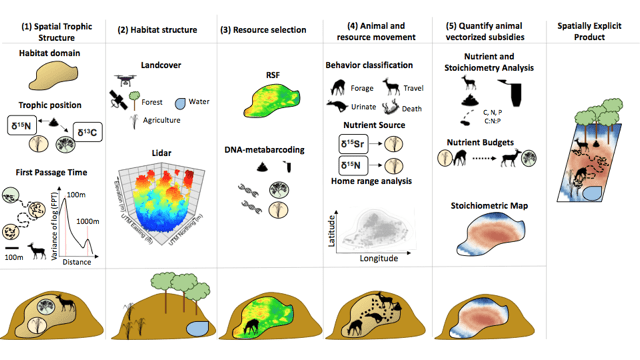
Our paper seeks to address just this challenge; providing a roadmap to guide all different animal-mediated nutrient translocation studies. We begin by providing a bit of theoretical background on meta-ecosystem theory to help guide question development. We then break down the structure of zoogeochemical studies in five steps. The first step is to address how, why, and where an animal is moving. Recently, technological advances in remote sensing and biologging have allowed us to begin to peek into the world of animal movement, providing a deep understanding of the habitats a given animal occupies, selects and avoids. We provide detailed guidance on various technologies to help researchers identify their study species movement. Next, the habitats used by the animals of interest must be identified and characterized, particularly the source and recipient habitats of nutrient inputs. We discuss the power of integrating tools such as LiDAR, topographic products, and other remotely sensed environmental products to help achieve a deep understanding of the habitats in question. In the third step, the nutrients available to, and consumed by, the animal must be identified and characterized in both the source and recipient habits. We discuss this can be achieved by tying various spatial ecology tools with dietary and trophic analyses. With this information we can take the fourth step; identifying the movement rate and directional flows of the species as well as any animal-vectorized subsidies. Many tools across remote sensing, biogeochemistry and spatial ecology can be used to tackle this question, and we discuss how the integration of these tools lead to clearer understanding. Finally, using all the above information, along with classic biogeochemical methods, we can identify both the quantity and location of nutrient deposition by the animal in a spatially explicit manner.
Together, these five steps provide a comprehensive picture of the nutrient subsidies animals move. Leaning on our backgrounds, study systems, and expertise we then walk through case studies to illustrate how using these five steps can create comprehensive studies and insights of animal-mediated nutrient translocations.
The five steps we outline in our paper are all extremely important to get a full picture of how animals move nutrients and connect ecosystems, but each step relies on tools sets developed in the disparate disciplines of ecosystem ecology, animal ecology, and remote sensing. However, when used together, these toolboxes can lead to integrative, coherent understandings of how animal vectored subsidies drive spatial ecosystem structure and functioning. Though providing a roadmap, we hope to help those embarking on zoogeochemistry studies to utilize a variety of the tools available to them.
Citations:
Julia E., & Zollner, P.A. Advancing research on animal‐transported subsidies by integrating animal movement and ecosystem modelling (2017) Journal of Animal Ecology 86.5: 987-997.
Roman, J. & McMarthy J.J. The whale pump: marine mammals enhance primary productivity in a coastal basin. (2010) PloS one 5.10: e13255
Schmitz, O.J. et al. Animals and the zoogeochemistry of the carbon cycle (2018) Science 362.6419 .
Wilcove, D.S., Wikelski, M. (2008) Going, going, gone: is animal migration disappearing. PLoS Biology 6.7: e188.
Read the paper
Read the full paper here: Ellis-Soto, D., Ferraro, K.M., Rizzuto, M., Briggs, E., Monk, J.D. and Schmitz, O.J. (2021), A methodological roadmap to quantify animal-vectored spatial ecosystem subsidies. J Anim Ecol. Accepted Author Manuscript. https://doi.org/10.1111/1365-2656.13538




One thought on “Animals in the driver’s seat: a methodological roadmap to animal-mediated nutrient translocation”