In the latest issue of the journal we have a new ‘How to…’ paper lead by Nick Fountain‐Jones from the University of Tasmania on How to make more from exposure data? An integrated machine learning pipeline to predict pathogen exposure. In this blog, Nick goes beyond the paper and discusses 5 mistakes and things to look out for when not only running the pipeline presented in the ‘How to…’ paper but when running any machine learning analytical routine.
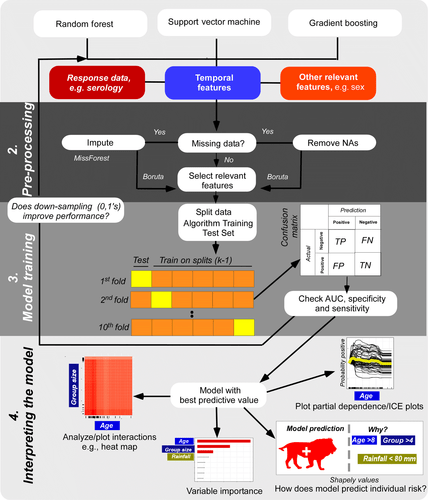
Statistical machine learning is now becoming much more commonplace in ecology. This is not surprising given the power of these methods to form robust predictive models from increasingly common big complex datasets. However, there is still lots of confusion about machine learning approaches and how they can be applied and interpreted. Our pipeline summarizes some of the latest advances in machine learning and puts them together in a way that can be used to help address not only exposure risk but a variety of ecological problems. We have tried to make our pipeline user-friendly and make it difficult to go too far wrong, however, mistakes happen and these can impact results. To avoid some of these pitfalls we list 5 mistakes and things to look out for when not only for running our pipeline but when running any machine learning analytical routine.
- The algorithm is only as good as the data
This is an obvious one, but always a good reminder. To politely put it, if you are putting rubbish data into a model, the output of the model no matter how powerful the algorithm, will be rubbish. Even though a random forest, for example, can run with thousands of predictors (or in computer science speak ‘features’) doesn’t mean you should add thousands of predictors. Think about what is reasonable given your hypotheses. Data visualization is also key and can help find mistakes and/or rubbish data. Recently we’ve been using the R package DataExplorer to streamline this process.
On a related note, missing data is common in ecological datasets and how you deal with it is important and worth its own blog article. In our paper, we provide a quick overview and recommend the ‘missForests’ R package if you go down the imputation route. Whilst this is a powerful method and works surprisingly well even on categorical data, always be cautious and check the performance of the algorithm carefully (normalized root mean square error [NRMSE] for continuous variables or proportion of falsely classified entries [PFC] for categorical data).
- It’s all about balance
Speaking of bad data, if you are trying to build a model and your binary response data is 95% zeros/ones or negatives/positives, then there are going to be problems. Machine learning models are lazy and will just predict the negative or positive class, i.e., the prediction will always be that the species won’t be there. ‘Class imbalance’ is often overlooked but can be an important problem. Even a minor class imbalance can lead to biased predictions, and we suggest always comparing your non-balanced dataset to a more balanced one. We provide some code that makes down-sampling your dataset easy but resampling or up-sampling (sampling with replacement) can be an option too.
- Always learn on a small dataset and try a simple model first
If you have a large dataset that takes a while (hours, days or weeks) to run on your machine/server make sure you do a trial run on a subset of the data to make sure everything is running OK. This is particularly important if you aren’t familiar with our pipeline or machine learning in general. There is nothing worse than waiting a long time to find out there is an error that must be fixed somewhere.
Consider always running a simple model first like a linear regression. I’m guilty of not doing this in the past when I know there are going to be complex interactions, but there are really no excuses as these models are the most interpretable. The R package ‘caret’ (Classification And REgression Training) that is at the heart of our pipeline can, without much recoding, allow you to compare the performance of your simple linear model with a more complex support vector machine for example. We show how to do this in our vignette. More complex may not mean better.
- Pay attention to model tuning
Tuning a machine learning model used to be a pain – manually going through different parameters and seeing how they impact the performance of the algorithm was time-consuming. The caret package has streamlined this process with tuning grids which enable many combinations of tuning parameters to be tested automatically with the best one used in the final model. Each algorithm has its own tuning parameters and these can be found on the caret GitHub page which we highly recommend: https://topepo.github.io/caret. You can also find them by typing ‘modelLookup(model=’gbm’)’. Whilst we have provided standard tuning grids for the three algorithms we compare in our paper, they may not be sufficient in some cases. It may be worth using tuneLength that picks a user given number of parameters to go through to check if any improvements in performance can be made. Always check the resampling profile of your model as these are key to visualizing how your tuning parameters are performing. Just use ‘plot (model)’ to do this.
- Rerun and interrogate models.
These models are often stochastic so it is always a good idea to check for consistency across runs. Use the same starting point (the seed number) for each model to give the same sequence of random numbers and report the seed you used to make the model reproducible. If you are getting strongly differing results with different seeds, creating a voting ensemble with an R package such as caretEnsemble is good practice.
Lastly, we provide a useful set of tools to interpret these complex models and using them will provide insights into how well your model is performing. In particular, calculating Shapely values will help work out why, for example, your model misclassified an observation and provide interesting insights into the mechanics underlying your model.
